MAGeCK is a computational tool to identify important genes from the recent genome-scale CRISPR-Cas9 knockout screens (or GeCKO) technology. MAGeCK can be used for prioritizing single-guide RNAs, genes and pathways in genome-scale CRISPR/Cas9 knockout screens. MAGeCK identifies both positively and negatively selected genes simultaneously and reports robust results across different experimental conditions. MAGeCK is developed and maintained by Wei Li and Han Xu from Prof. Xiaole Shirley Liu’s lab at the Department of Biostatistics and Computational Biology, Dana-Farber Cancer Institute and Harvard School of Public Health. MAGeCK has been used to identify functional lncRNAs from screens with close to 100% validation rate.Documentation Index
Fetch the complete documentation index at: https://wiki.latch.bio/llms.txt
Use this file to discover all available pages before exploring further.
How to run MAGeCK on Latch
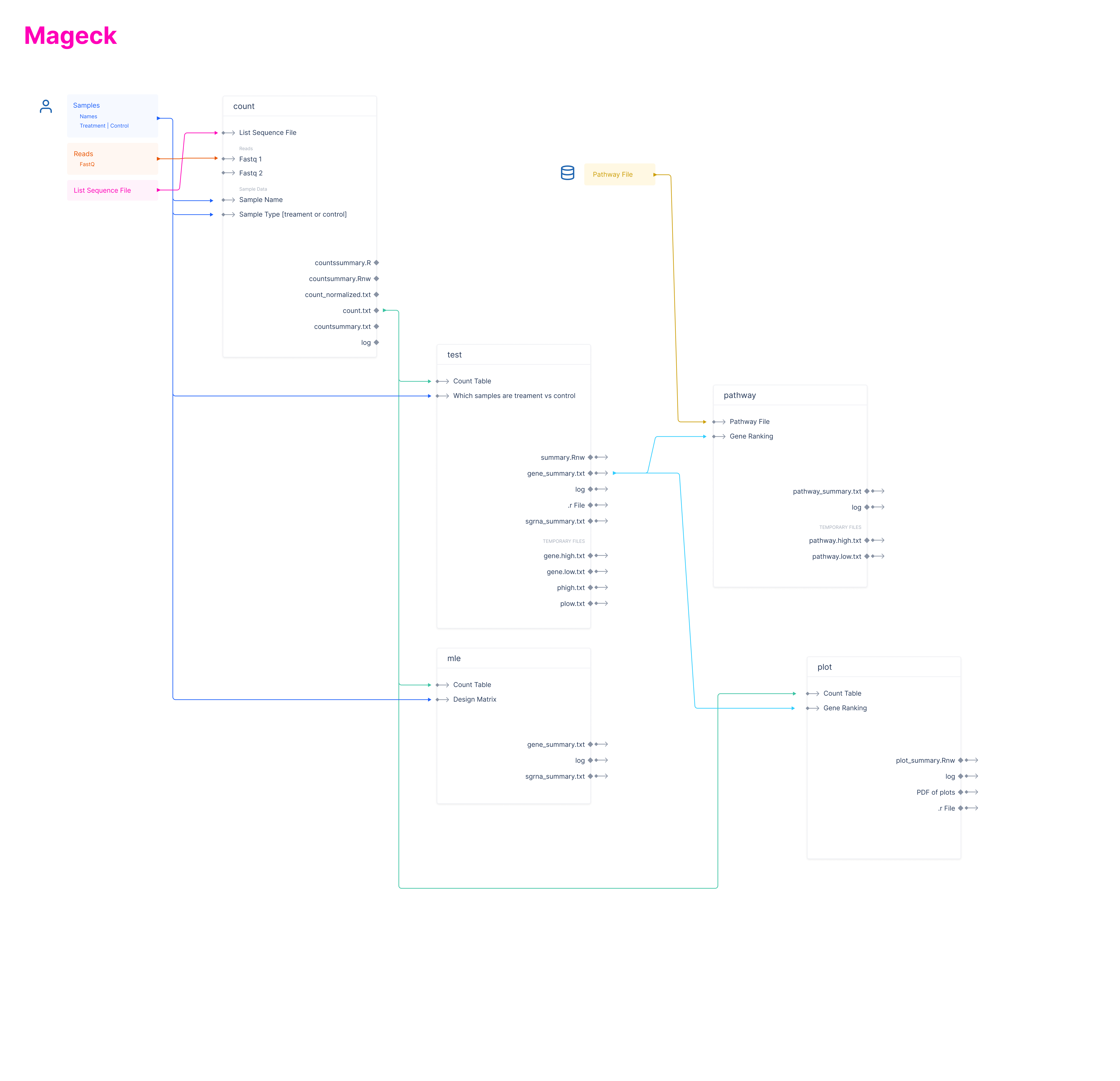
On Latch we have split up the MAGeCK workflow into its subcommands to be run. These are:MAGeCK - Count
MAGeCK - Test
MAGeCK - MLE
MAGeCK - Pathway
MAGeCK - Plot
mageck-subcommand-input-output-graph.pdf
1284.5 KB